Migraine distribution page
Migraine implements coalescent algorithms for maximum likelihood analysis of population genetic data. It considers Allelic counts and DNA sequences. The code is designed to be compiled with the GNU C++ compiler under Windows and Linux (including its Mac OSX incarnation).
Recent changes (from version 0.5 upwards): Migraine now includes the Infinite Site Mutation (ISM) model, a new demographic model, OnePopFounderFlush, and a resampling procedure improving the importance sampling algorithm in the OnePopVarSize model. Further, it now uses a standard R package, blackbox, rather than the older Rmigraine package that was previously distributed through this page, so that installation is easier.
The different models that can be fitted
are:
- one- and two-dimensional isolation by distance models (including the island model as a subcase) (Rousset and Leblois 2007,2012);
- the two-populations model described in de Iorio et al. (2005);
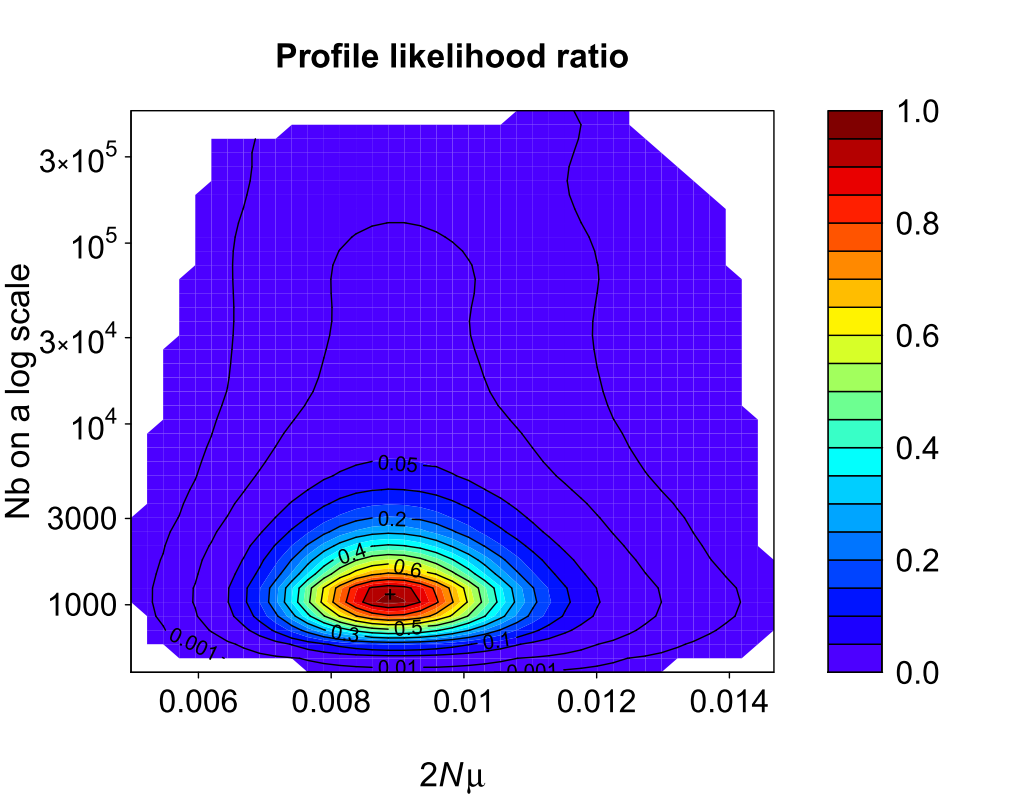
- the elementary one-population model (with 2Nμ as unique parameter);
- the OnePopVarSize model, including markers evolving under a GSM (Leblois et al., 2014) or an ISM;
- the OnePopFounderFlush model.
Typical analyses can be performed in seconds (in the one-parameter case) or overnight (in some three-parameter cases), and longer analyses can easily been split among different computers.
There are two versions of the documentation, a short one that should be sufficient for most applications, and a longer one for complications, which hopefully no one needs to read.
For Windows executable (migraine.exe) in 64-bit environments, download migraineWin64.tar.gz.
For the sources for all operating systems, download migraine.tar.gz (for helper Nexus2GP program, download sourcesNexus2GP.tar.gz).
For the "first session" sample files, download examples.zip.
See the documentation for all further information.
Migraine (C) François Rousset & Raphaël Leblois 2007-present, with additional contributions by Champak R.Beeravolu 2013-2014.
Migraine is freeware (i.e. you don't need to pay). It is free software covered by the CeCILL licence (GPL compatible), i.e. it can be used, copied, studied, modified, and redistributed in other free software (i.e. also covered by a GPL-compatible licence, with freely available source code, even if commercial software) provided the Migraine source is acknowledged.
This page (C) F. Rousset 2007-present
C++ sources/executables last
updated on 2020/10/01